intrinsically disordered proteins
explore IDP structural heterogeneity
Intrinsically disordered proteins (IDPs) are a fascinating class of proteins that are characterized by their lack of a fixed, ordered structure. Unlike the classical paradigm of protein folding, IDPs exhibit high structural flexibility, allowing them to interact with multiple binding partners and assume a variety of conformations. This heterogeneity of their structural ensemble is a fundamental characteristic of IDPs and is essential for their diverse range of functions in cellular processes such as signaling, regulation, and molecular recognition. Indeed, many IDPs are involved in disease processes, and their dysfunction has been implicated in a range of disorders, including neurodegenerative diseases, cancer, and viral infections.
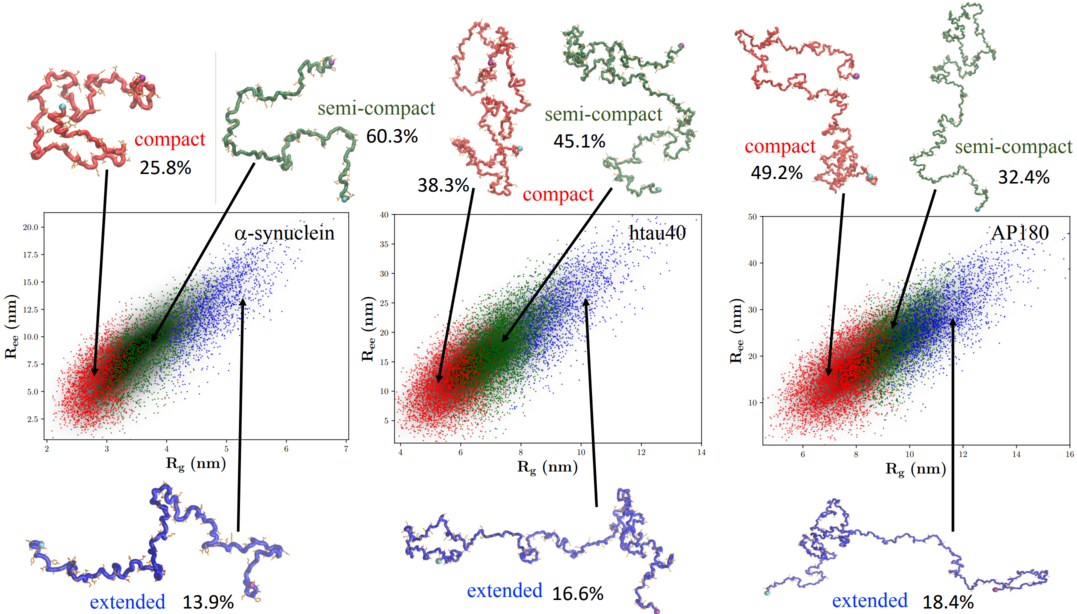
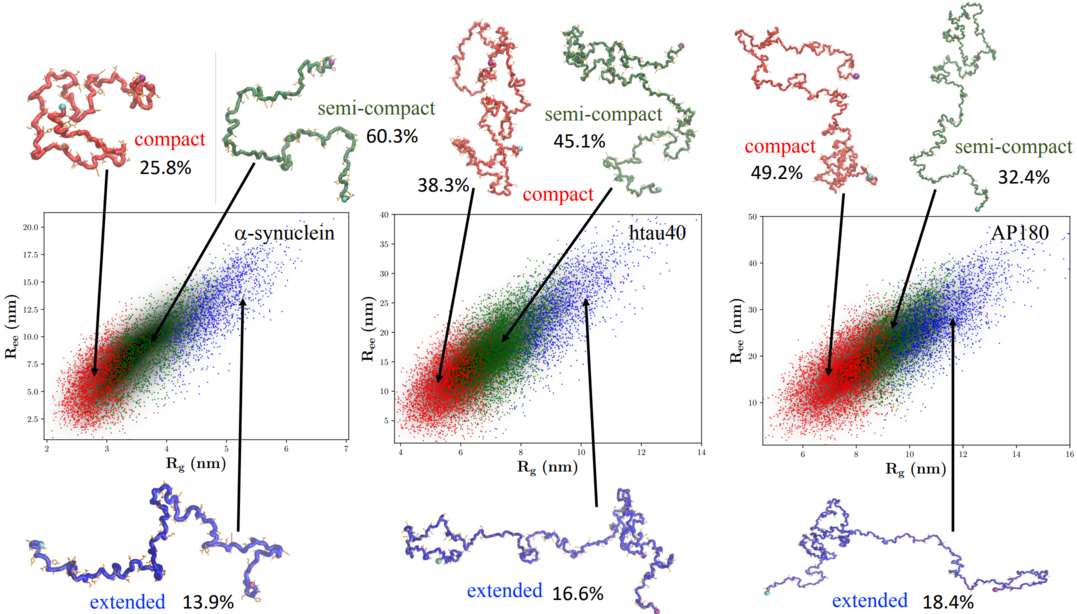
We have developed a computational model to quantitatively simulate IDP structural ensemble. Our simulations reproduced in excellent quality many structural aspects–including SAXS profiles (and thus the radius of gyration Rg), and NMR paramagnetic relaxation enhancement–of a diverse set of IDPs of different lengths and sequences. Simulations revealed that IDPs in general behave globally like self-avoiding random walk polymers, and the sequence-dependent features only emerge by careful analyses of the ensemble.